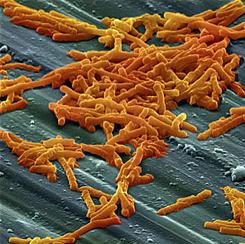
Clostridium difficile, more commonly referred to as C. diff, is a bacteria that makes half a million American's sick each year, and is responsible for over 25,000 deaths annually, both directly and indirectly. The bacteria can lead to serious illnesses in the gut, that can cause diarrhea and colon inflammation. Often times, C. diff infections can be caused by the over use of antibiotics, which affect the healthy bacteria in the gut and provide opportunity for C. diff bacteria to grow in that area.
Researchers from the University of Michigan, Ann Arbor have received a five-year, $9.2 million grant from the National Institute of Allergy and Infectious Diseases to further study C. diff, to learn more about it with the aim of developing new treatment methods. (Image of C. diff bacteria courtesy of Cjc2nd via Wikimedia Commons)
University of Michigan researcher and professor Dr. Vincent Young explained that this grant "should help us lay the groundwork for both short- and long-term solutions to the C. difficile crisis, by understanding what puts patients at risk for C. difficile infection, and how we can best protect and cure them."
The Ann Arbor researchers will be creating computational models to track and better understand the path that the C. diff bacteria take when infecting the gut, in order for them to be able to predict its movement. The team will create these models using data collected both in their lab and from blood samples and patient records from actual patients with C. diff. The data collected in these models will provide researchers with information about how the natural microbe population in the gut, a patient's health, and antibiotics all contribute to the cause of C. diff infections.
Through previous research with mice, the University of Michigan researchers have already detailed the life cycle of C. diff in the gut, which has helped them to understand how normal gut bacteria interferes with the C. diff bacteria that can appear in the gut. This knowledge from both humans and mice will help the researchers better understand what risk factors contribute to a patient's vulnerability to C. diff infections.
This $9.2 million grant is part of a $1.2 billion effort by the United States government to research and defeat bacteria that are resistant to antibiotics.



(Images via Biotechnology Calendar, Inc., Wikimedia Commons, Biotechnology Calendar, Inc. respectively)
The University of Michigan, Ann Arbor is a leading institution in terms of the amount of funding it receives annually and research it produces. In the 2015 fiscal year, the university received more than $456 million in funding from the National Institutes of Health (NIH). This funding will greatly benefit multiple research projects and new building constructions, including:
-
RELATED NEWS:
UM Professor Tells Cells to Regrow Bone
Portland Scientists Awarded $3M for Tuberculosis Vaccine Research
A $261 million project was approved to construct a new teaching, research, and museum facility for the biological sciences. The 300,000-square-foot building will provide modern, world-class facilities for UM’s biological science programs and will be completed in 2019.
-
NIH grants totaling $6.3 million renewed funding for continued research projects at The University of Michigan Comprehensive Cancer Center.
-
The University of Michigan was awarded a $2.4 million NIH grant to investigate the role of epigenetics in disease risk.
With all this funding, researchers at the University of Michigan, Ann Arbor have the means to purchase many new lab products that will help with their studies and clinical trials. Biotechnology Calendar, Inc. produces an annual BioResearch Product Faire™ Event in Ann Arbor that is a great opportunity to market lab products to active life science researchers at the university. This annual event brings active researchers together with scientific supply companies, so that the researchers can find the best and newest products and technologies to further their work.
The 16th annual BioResearch Product Faire™ at UMich will be held on July 28, 2016, where more than 500 life scientists are expected to attend. Last year, 546 researchers came from 38 different research buildings and 86 on-campus departments.
To learn more about participating in this popular event, visit the link below:


